 Any other source of DNA treated with the same enzyme will produce such molecules. These are called "sticky ends" because they are able to form base pairs with any DNA molecule that contains the complementary sticky end. The ends of the cut have an overhanging piece of single-stranded DNA. However, many restriction enzymes cut in an offset fashion. HaeIII and AluI cut straight across the double helix producing "blunt" ends. These fragments can be separated from one another and the sequence of each determined. Thus treatment of this DNA with the enzyme produces 11 fragments, each with a precise length and nucleotide sequence. This particular sequence occurs at 11 places in the circular DNA molecule of the virus φX174. The cut is made between the adjacent G and C. For example, the bacterium Hemophilus aegypticus produces an enzyme named HaeIII that cuts DNA wherever it encounters the sequenceģ'CCGG5' Figure 5.7.2: Restriction Enzymes Figure 5.7.1: Restriction DigestĪ restriction enzyme recognizes and cuts DNA only at a particular sequence of nucleotides. The rarer the site it recognizes, the smaller the number of pieces produced by a given restriction endonuclease. The tools for this are the restriction endonucleases. What is needed is a way to cleave the DNA molecule at a few precisely-located sites so that a small set of homogeneous fragments are produced. This produces a heterogeneous collection of fragments of varying sizes. Many DNA-digesting enzymes (like those in your pancreatic fluid) can do this, but most of them are no use for sequence work because they cut each molecule randomly. To be able to sequence DNA, it is first necessary to cut it into smaller fragments. Because they cut within the molecule, they are often called restriction endonucleases. Restriction enzymes are DNA-cutting enzymes found in bacteria (and harvested from them for use). NOT FOR USE IN DIAGNOSTIC PROCEDURES (EXCEPT AS SPECIFICALLY NOTED).\) Our mission is to develop high-quality innovative tools and services to accelerate discovery.įOR RESEARCH USE ONLY. As a member of the Takara Bio Group, Takara Bio USA is part of a company that holds a leadership position in the global market and is committed to improving the human condition through biotechnology. provides kits, reagents, instruments, and services that help researchers explore questions about gene discovery, regulation, and function. 
Compositions of provided universal buffers and other reagents Since BSA and Triton X-100 are supplied with enzymes at a 10X concentration (0.1%), be sure to add 1/10 the volume of the reaction mixture to the buffer before starting the reaction. Some restriction enzymes are supposed to exhibit 100% activity when BSA or Triton X-100 is added to the reaction system. Since 10X T4 buffer does not contain BSA, be sure to add BSA to a final concentration of 0.01%.  
Please add 1/10 the volume of the reaction mixture.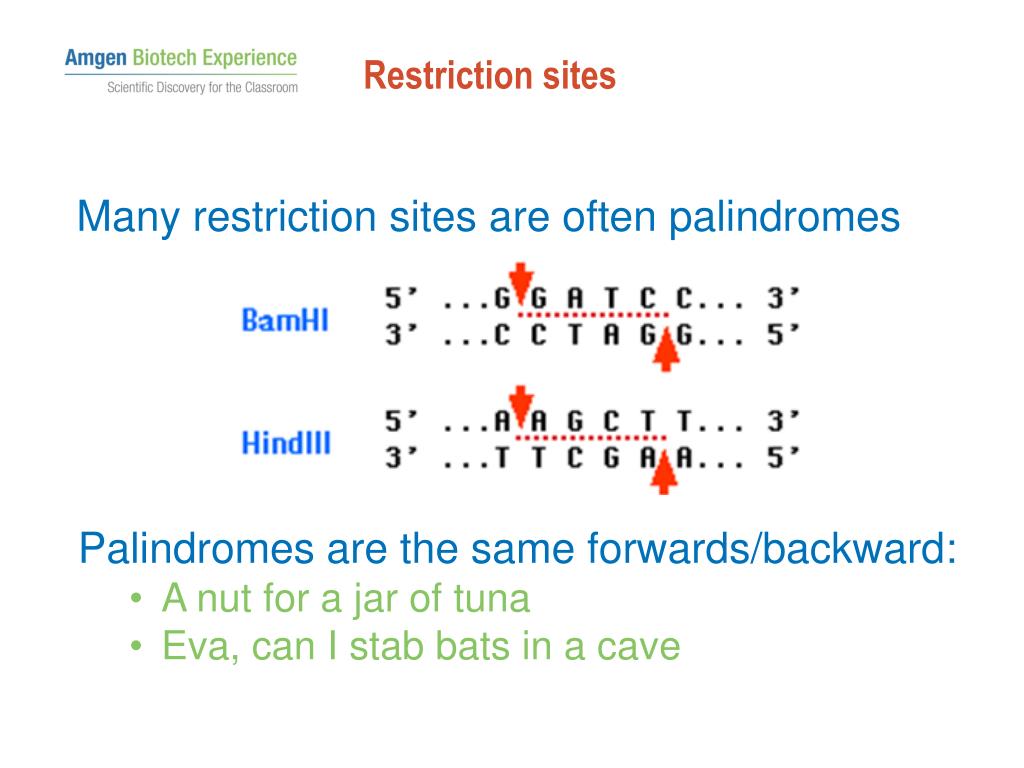 
Universal buffers are provided at a 10X concentration. Instructions for the use of universal buffers If there is a precipitate, dissolve in a warm water bath before use. Note: SDS may precipitate during storage at room temperature. The tables below list relative activities of restriction enzymes in the universal buffers and instructions for use and compositions of buffers.ġ/10 volume of 10X loading buffer to stop the digestion reaction, and run on an agarose gel. The compositions of these basal buffers vary depending on the enzyme.įor information on double digestion, see the universal buffers for double digestion with restriction enzymes page. In order to avoid these effects, use of buffers highlighted in blue or pink is recommended.Ī few specific enzymes (AccIII, BalI, BcnI, BglI, Bpu1102I, Cfr10I, Eco52I, NruI, PshBI, SnaBI, SspI, TaqI, and VpaK11BI), are each supplied with a basal buffer specialized for the particular enzyme. Values in ( ) indicate buffers that are likely to be affected by star activity. The relative activity in each of the other universal buffers is normalized to the optimal buffer, where the activity of each enzyme in the optimal buffer is expressed as 100%. Our restriction enzymes are supplied with an optimal universal buffer (one of five universal buffers indicated in blue in the table below). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
